Unravelling Enzymatic Reaction Mechanisms
We investigate the operation of protein engines that convert chemical energy into motion.
Read about our work here: Nature Chemistry 2024, J. Chem. Theory Comput. 2024, J. Am. Chem. Soc. 2020, Biochem. Soc. Trans. 2021, Proc Natl. Acad. Sci. U.S.A 2017, and eLife 2016.
The catalysis of the ATP-hydrolysis reaction and how it couples energy release to motion is a long-standing topic in enzymology research. Experimentally grounded snapshots of short-lived reaction intermediates are crucial to substantiate the stepwise dissection of enzymatic reaction cycles. However, for structural investigations, enzymes are frequently modified or inhibited to trap uniform, static states, increasing the potential for distorting the authentic molecular mechanisms.
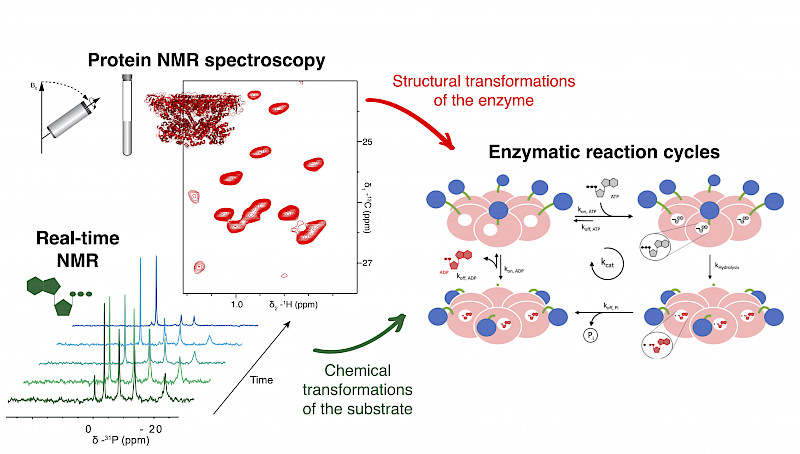
Real-time NMR spectroscopy can bridge the gap between static enzyme structures and the dynamics of substrate conversion.
Many functional enzyme complexes reach molecular sizes that exceed the limits of solution-state NMR. We make the such complexes amenable to magic angle spinning solid-state NMR via sedimentation techniques. To this end, we have established an integrated approach that permits to monitor simultaneously conformational transitions of substrate and enzyme.
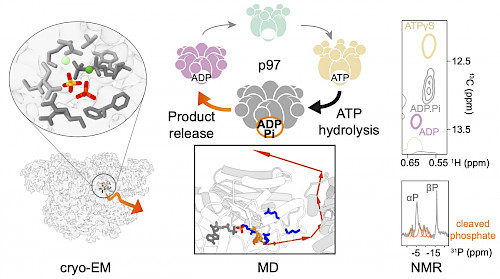
We have joined forces with collaborators from in cryo-electron microscopy and computational chemists to overcome present limits in the characterization of short-lived and heterogeneous reaction intermediates. Our goal is to develop a comprehensive picture of the structural dynamics that accompany the catalytic reaction cycle in different types of NTPases.
We are particularly intrigued by the p97/Cdc48 AAA+ protein, which fuels vital processes in the human cell, where it represents one of the most abundant soluble proteins. p97 is an intensely explored pharmacological target in the therapy of cancer and degenerative diseases. A deeper mechanistic understanding is highly relevant to enzyme design and drug discovery, not only of this prototype member of the AAA+ ATPase family, but also of the broader P-loop NTPases.